Introduction to monogram
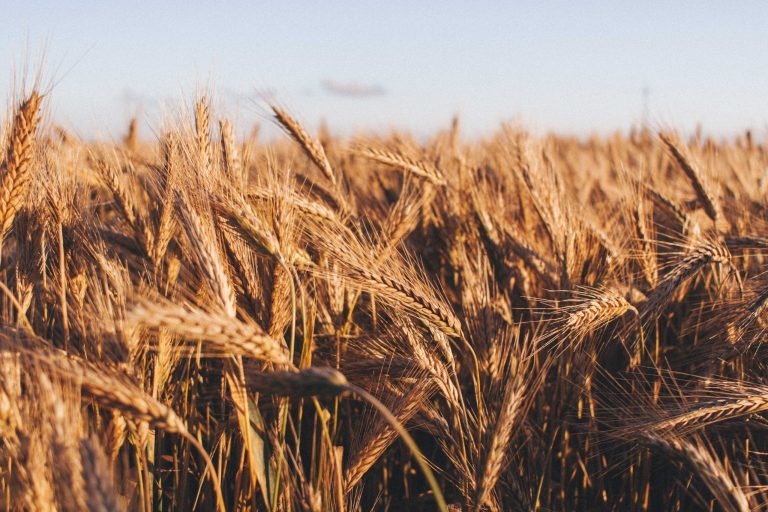
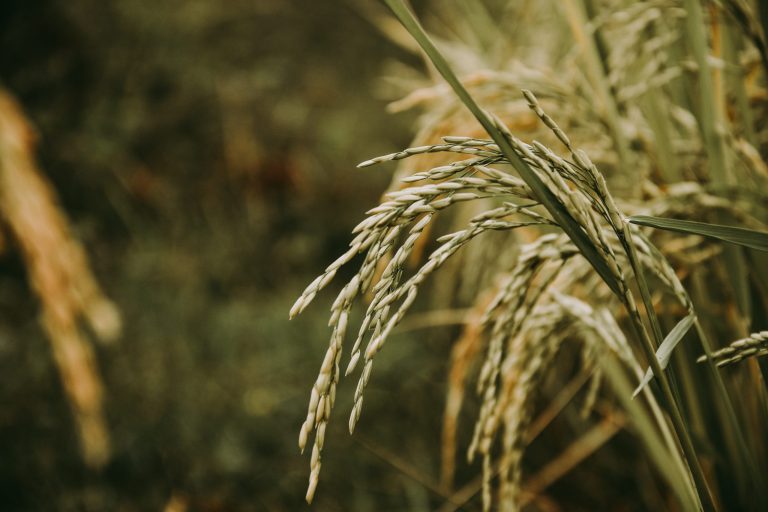
The Monogram Network consists of UK based researchers with an active interest in small grain cereal and grass (including the C4 energy grasses) research.

about us
The Monogram Network consists of UK based researchers with an active interest in small grain cereal and grass (including the C4 energy grasses) research. Commercial scientists and plant breeders are active members and provide the link between Monogram science and commercial exploitation. Monogram includes both basic and more applied research and its members span disciplines including plant genetics, physiology, pathology, breeding, and bioinformatics. We also have extensive expertise in outreach activities and links with industry.
Resources
We have a list of resources available for multiple different species and purposes
Our Latest News
Solving the living puzzle: Homoeolog interaction in polyploid genomes
Many economically important crops, such as wheat, cotton and coffee are allopolyploids, i.e., they originated from hybridisation of two or more species followed/preceded by whole

My Journey in Wheat
Ever since I was a teenager, I have been inspired by the “Golden Rice” story and amazed by how the application of plant sciences can

Unravelling the genetic relationships between cereals and their microbiome
Cereals, like other plants, live in association with a myriad of microorganisms collectively referred to as the plant microbiome. These interactions are particularly dynamic in




